在复现 GitHub 上的 SMT_Tutorial 代码时,发现生成的 3D 图片中点数出现畸形。请问如何解决这个问题?
在复现 GitHub 上的 SMT_Tutorial 项目时,2D 图片能够正常生成,但 3D 图片在复现时出现了点数畸形,并且呈现竖条状分布。请问这可能是什么原因,如何解决这个问题?
from future import print_function, division
import numpy as np
from scipy import linalg
from smt.utils.misc import compute_rms_error
from smt.problems import Sphere, NdimRobotArm, Rosenbrock
from smt.sampling_methods import LHS
from smt.surrogate_models import LS, QP, KPLS, KRG, KPLSK, GEKPLS, MGP
#to ignore warning messages
import warnings
warnings.filterwarnings(“ignore”)
try:
from smt.surrogate_models import IDW, RBF, RMTC, RMTB
compiled_available = True
except:
compiled_available = False
try:
import matplotlib.pyplot as plt
plot_status = True
except:
plot_status = False
import scipy.interpolate
from mpl_toolkits.mplot3d import Axes3D
import matplotlib.pyplot as plt
from matplotlib import cm
########### Initialization of the problem, construction of the training and validation points
ndim = 2
ndoe = 20 #int(10*ndim)
Define the function
fun = Rosenbrock(ndim=ndim)
Construction of the DOE
in order to have the always same LHS points, random_state=1
sampling = LHS(xlimits=fun.xlimits, criterion=’ese’, random_state=1)
xt = sampling(ndoe)
Compute the outputs
yt = fun(xt)
Construction of the validation points
ntest = 200 #500
sampling = LHS(xlimits=fun.xlimits, criterion=’ese’, random_state=1)
xtest = sampling(ntest)
ytest = fun(xtest)
#To visualize the DOE points
fig = plt.figure(figsize=(10, 10))
plt.scatter(xt[:,0],xt[:,1],marker = ‘x’,c=’b’,s=200,label=’Training points’)
plt.scatter(xtest[:,0],xtest[:,1],marker = ‘.’,c=’k’, s=200, label=’Validation points’)
plt.title(‘DOE’)
plt.xlabel(‘x1’)
plt.ylabel(‘x2’)
plt.legend()
plt.show()
To plot the Rosenbrock function
x = np.linspace(-2,2,50)
res = []
for x0 in x:
for x1 in x:
res.append(fun(np.array([[x0,x1]])))
res = np.array(res)
res = res.reshape((50,50)).T
X,Y = np.meshgrid(x,x)
fig = plt.figure(figsize=(15, 10))
ax = fig.add_subplot(projection=’3d’)
surf = ax.plot_surface(X, Y, res, cmap=cm.viridis,
linewidth=0, antialiased=False,alpha=0.5)
ax.scatter(xt[:,0],xt[:,1],yt,zdir=’z’,marker = ‘x’,c=’b’,s=200,label=’Training point’)
ax.scatter(xtest[:,0],xtest[:,1],ytest,zdir=’z’,marker = ‘.’,c=’k’,s=200,label=’Validation point’)
plt.title(‘Rosenbrock function’)
plt.xlabel(‘x1’)
plt.ylabel(‘x2’)
plt.legend()
plt.show()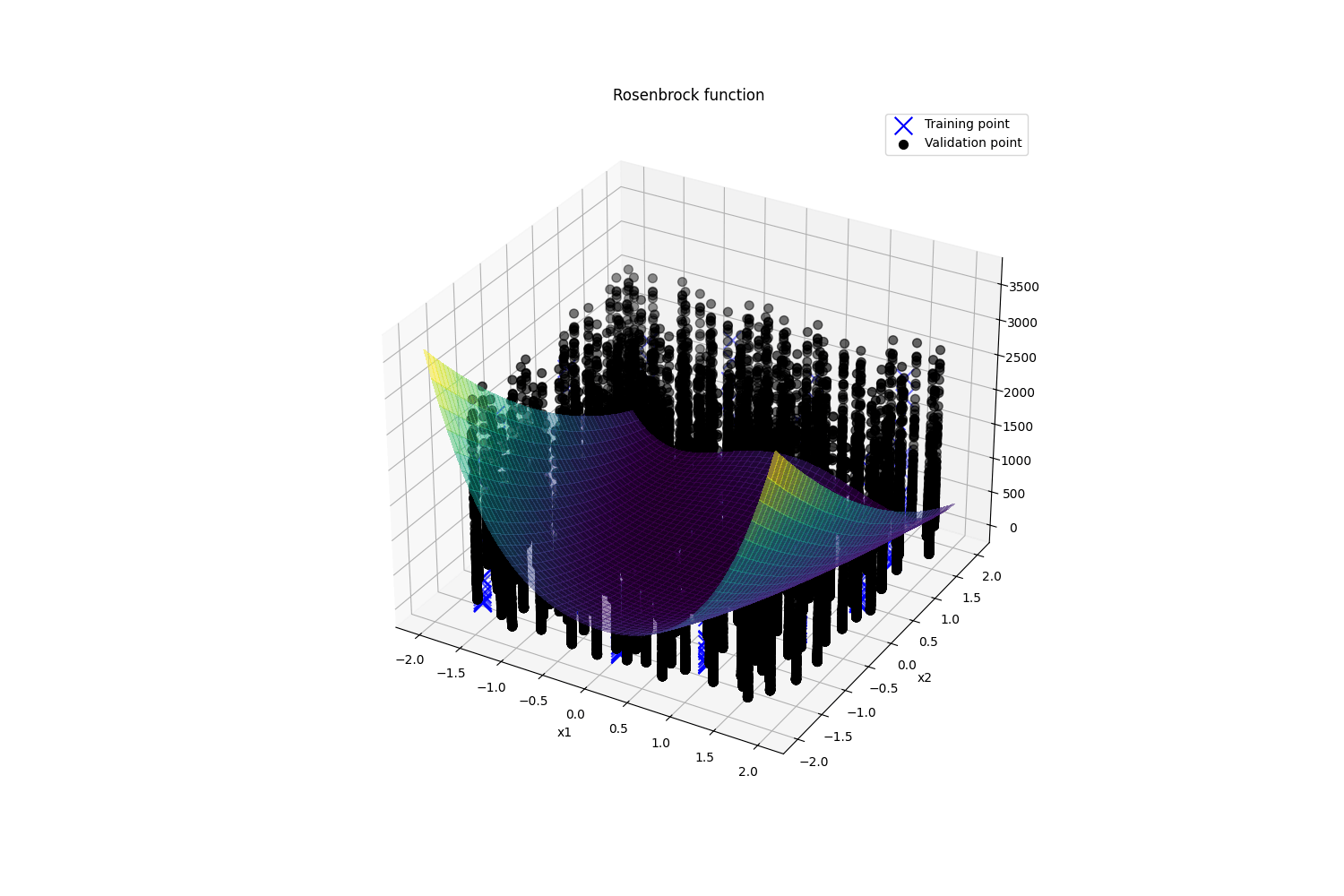
图1 复现代码畸形图片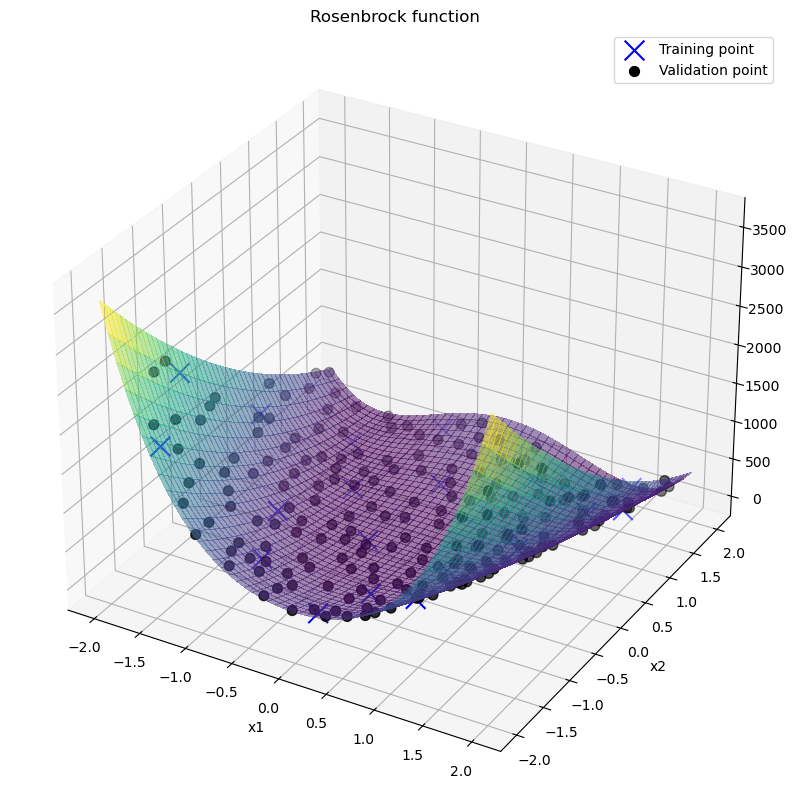
图2 项目作者生成图片




 关于 LearnKu
关于 LearnKu




推荐文章: